Research Groups
Group Ban
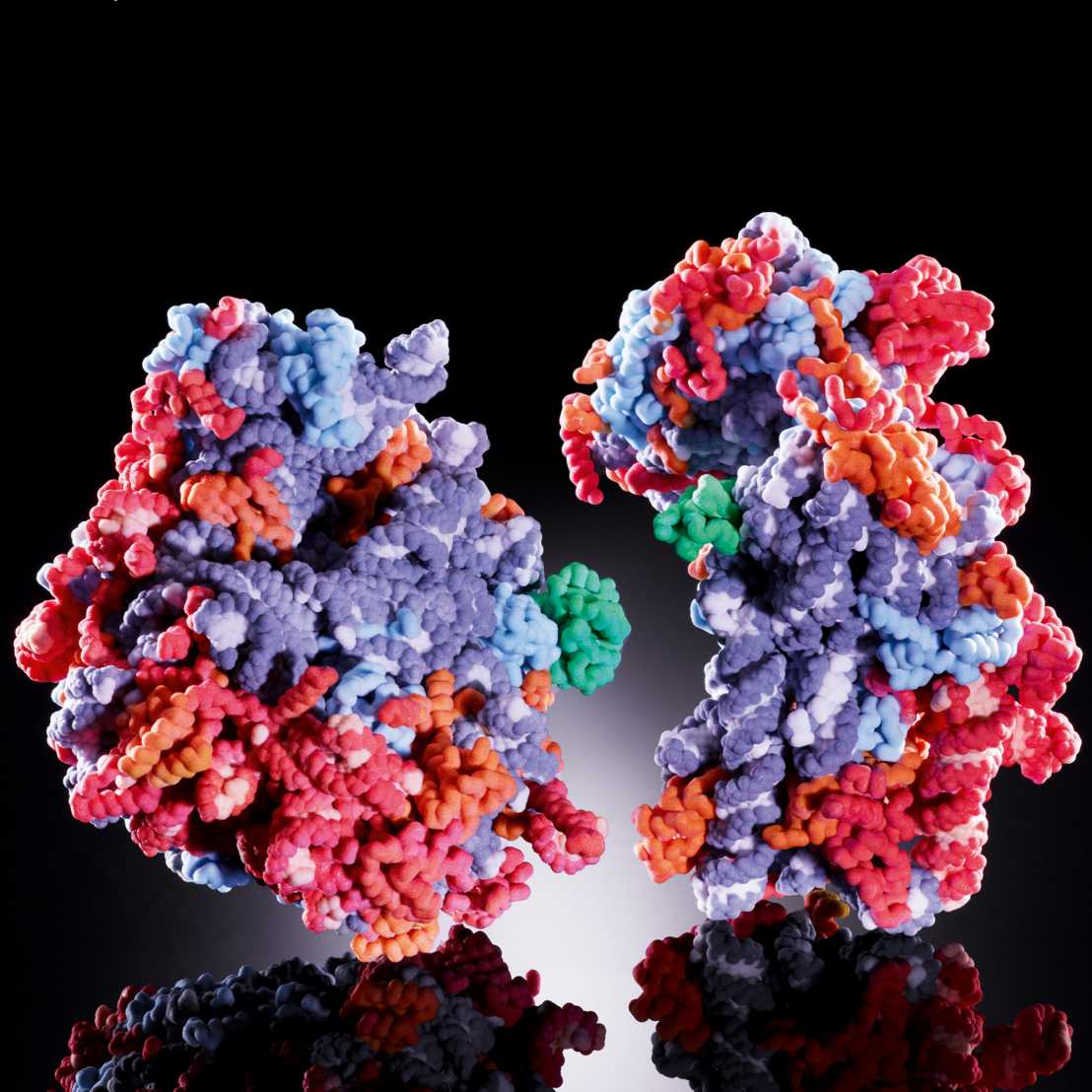
We are interested in understanding ribosomal structure and function in all kingdoms of life. Using a combination of biochemical, crystallographic and electron microscopic experiments we have obtained insights into co-translational nascent chain folding, processing and targeting to the membranes. Our recent focus is to understand eukaryotic cytoplasmic and mitochondrial ribosomes and their functional complexes involved in regulation and initiation of protein sythesis.
We carry out our experiments at the Swiss Light Source Synchrotron for crystal structure determinations and we use state-of-the-art cryo-electron microscopes for high-resolution structure determination by single particle reconstruction methods. Ultimately we intend to obtain a “molecular movie” of these processes at high resolution so that they can be understood at a detailed mechanistic level. Read more about our research.
Group Hospenthal
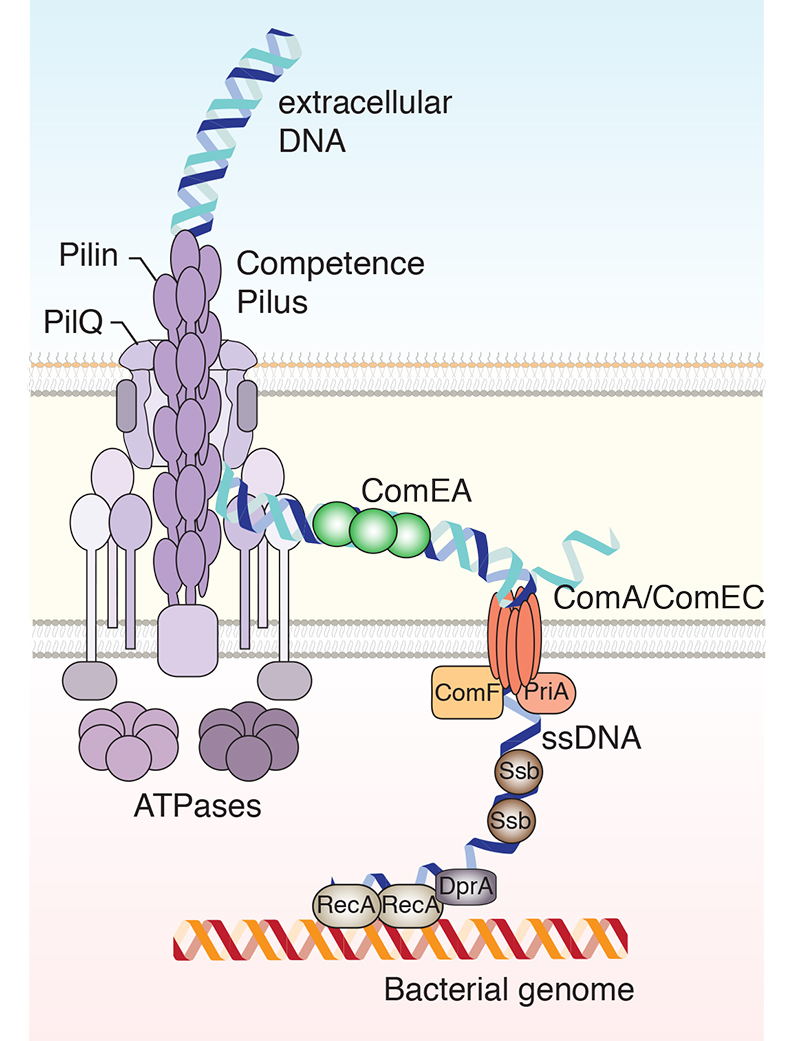
We seek to understand the processes that govern horizontal gene transfer (HGT) in bacteria. We are specifically interested in natural competence, one of the three main avenues of HGT. Naturally competent bacteria can undergo natural transformation by taking up free extracellular DNA and integrating it into their genome. This process contributes to the spread of antibiotic resistance and other pathogenicity traits amongst bacteria; an ever-growing problem for modern medicine. The overarching goal of our research is to understand how DNA uptake and translocation across the bacterial membranes is achieved. Our aim is to characterise the proteins involved biochemically, structurally and functionally. We combine cryo-electron microscopy (cryo-EM), X-ray crystallography and nuclear magnetic resonance (NMR) spectroscopy in a complementary manner.
Read more about our research.
Group Jonas
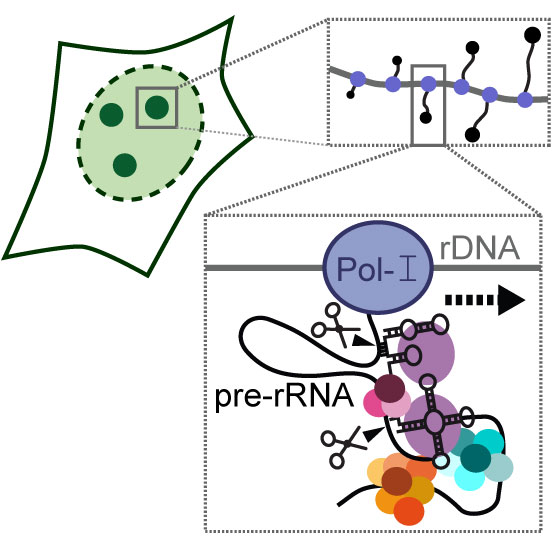
RNA-protein complexes (RNPs) have essential roles in many eukaryotic cellular processes from translation to splicing, protein translocation, RNA modification or telomere maintenance. Before they can carry out their functions these RNPs are assembled and matured themselves by dedicated machineries. We study the biogenesis processes of essential RNPs in the human nucleus at a molecular level. Our aim is to obtain a detailed mechanistic understanding of RNP assembly pathways, and the role of malfunctions in their assembly machineries during pathogenesis. Read more about our research.
Group Korkhov
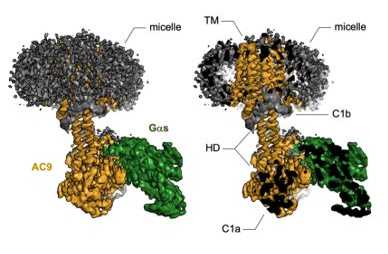
Interactions between the cells and their environment occur at the plasma membrane. Extracellular signals are sensed by plasma membrane-localized receptor proteins. These signals are then transmited across the membrane via multiple biomolecular interactions involving membrane-bound and soluble proteins, resulting in complex biochemical responses inside the cell. The process of signal transduction is critically important for physiology in health and disease. Our research is focused on understanding the molecular mechanisms of signal transduction. We use protein biochemistry, biophysics, X-ray crystallography and cryo-EM to gain a deeper understanding of the structure-function relationships of membrane proteins involved in various aspects of cellular signaling. Our current focus is on two major aspects of cellular signaling: the cAMP pathway (membrane adenylyl cyclases), and the Hedgehog signaling pathway (Patched and Dispatched). external page Read more about our research.
Group Locher
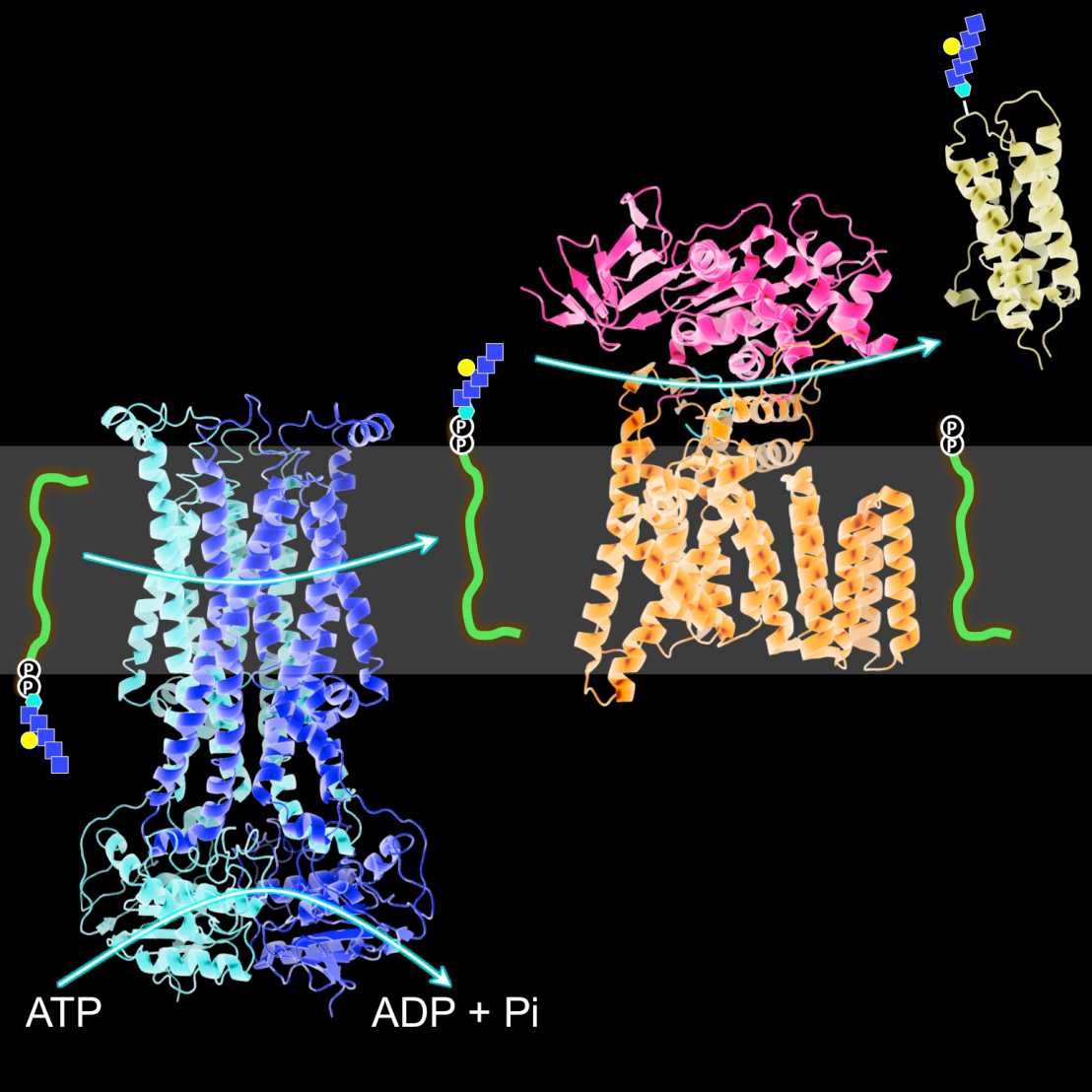
Transport of nutrients and waste products across membranes is essential to cellular life. We explore, at a molecular level, how integral membrane proteins transport substrates across the lipid bilayer. Our main focus is on ATP-binding cassette (ABC) transporters, a large family of proteins that couple unidirectional substrate transport to the hydrolysis of ATP. Human ABC proteins are associated with various diseases including cystic fibrosis or multi-drug resistance of cancer cells, whereas bacterial homologs mediate nutrient uptake and drug extrusion. We determine the structures of ABC transporters at high resolution using X-ray crystallography and we study their dynamics and in vitro transport kinetics after reconstitution in liposomes. Read more about our research.
Group Pilhofer
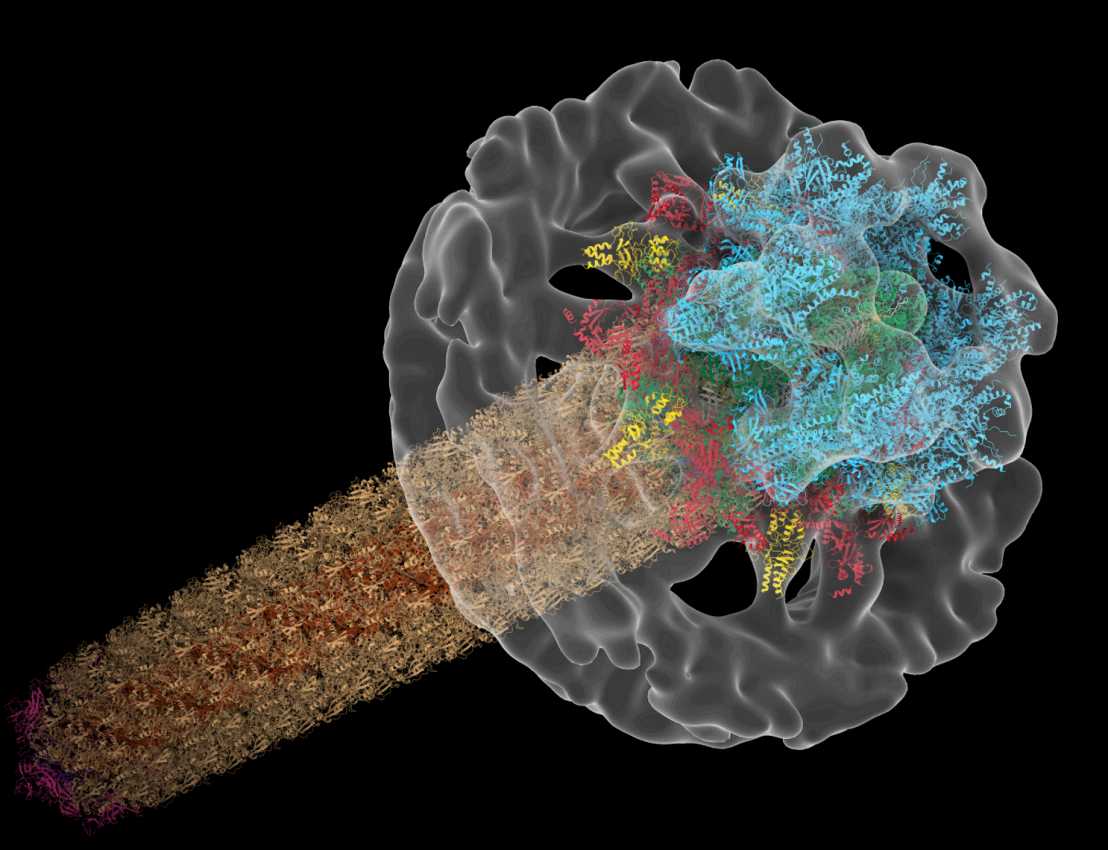
We are intrigued by the many ways bacteria interact with other cells. Our goal is to establish mechanistic models of macromolecular machines mediating these cell-cell interactions. We integrate information from the molecular to the cellular and intercellular scale, by combining our key technique cryo-electron tomography (cryoET) with other imaging techniques and functional assays. Read more about our research.
Group Weber-Ban
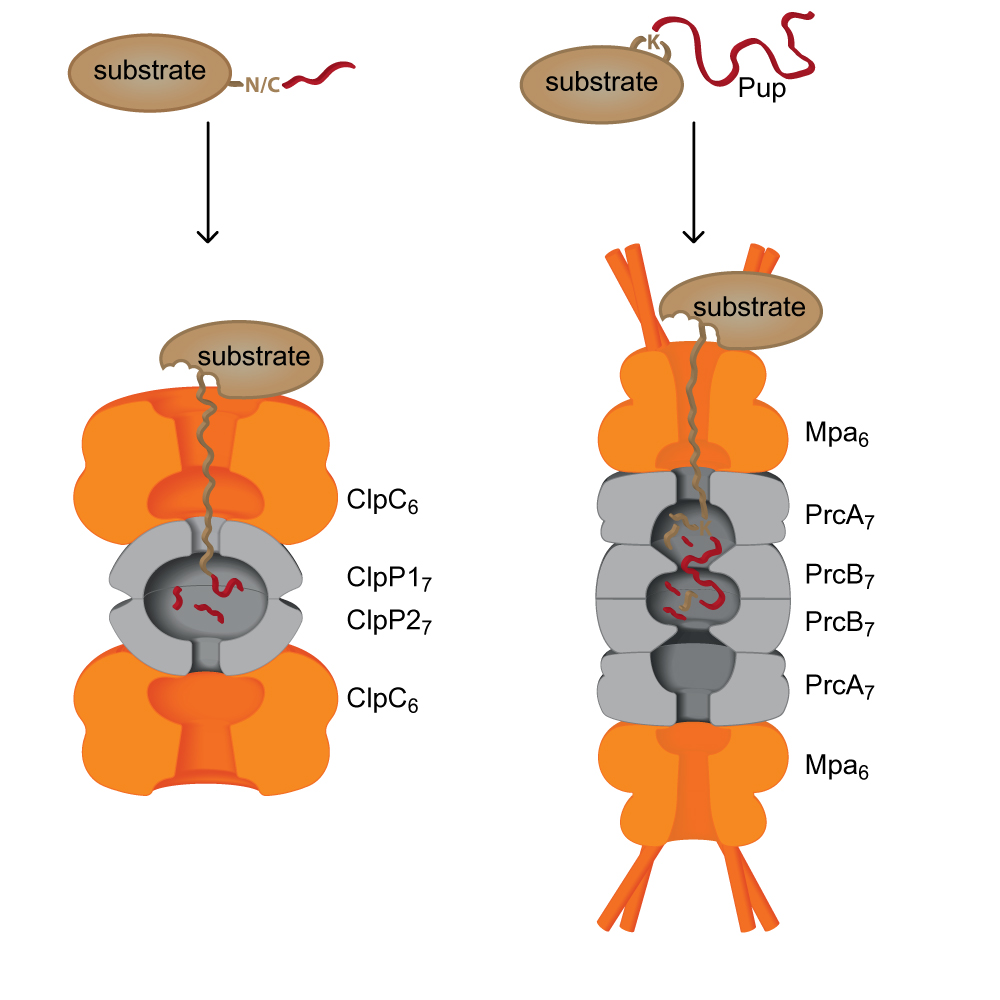
The Weber-Ban group is interested in chaperone-protease complexes, molecular degradation machines responsible for the processive breakdown of cellular proteins. In particular, we have recently focused our attention on a proteasome-system found in certain bacteria, amongst them the highly pathogenic Mycobacterium tuberculosis. Substrates are recruited by a ubiquitin-like pathway termed pupylation, wherein a small protein termed Pup (prokaryotic ubiquitin-like protein) is conjugated to lysine residues of protein substrates. Read more about our research.
Group Wieczorek
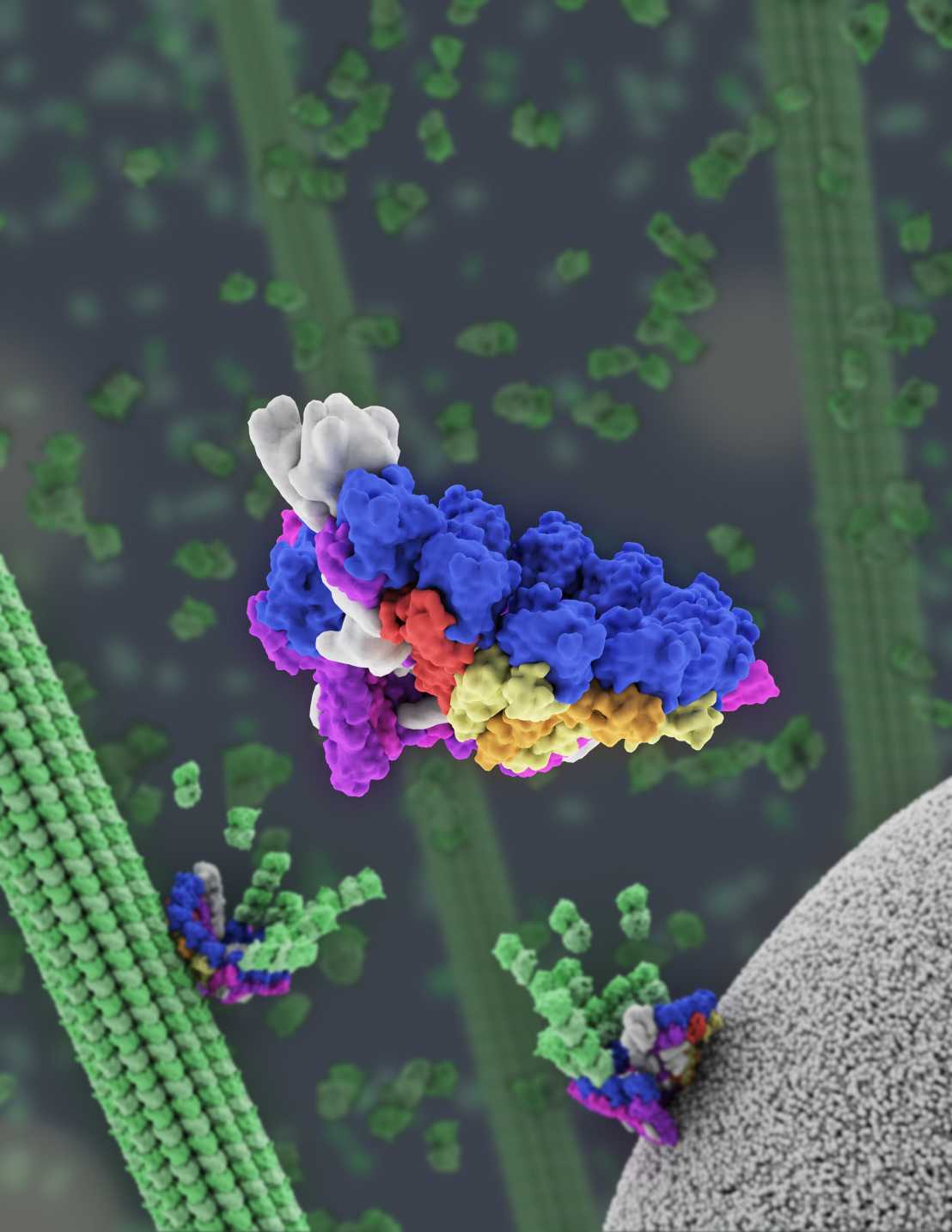
Our group is fascinated by the question of biological emergence: how do complex, self-regulating cellular assemblies arise from nanometer-scale components? To gain insight into this fundamental biological question, we use the microtubule cytoskeleton as a model system. A current focus of the lab is on the gamma-tubulin ring complex, a 2.3 MDa multiprotein assembly that nucleates microtubule growth and is essential for proper microtubule network organization in cells. The lab uses an interdisciplinary approach combining structural biology, biochemistry, single molecule microscopy, and cell biology. Read more about our research.
Group Wüthrich
The Wüthrich research group was initiated in 1969. Its research projects are in structural biology and protein research, with a specialization in nuclear magnetic resonance. The current research pursued in Zürich has been designed to extend and complement the structural biology data on prion proteins, which were accumulated during the last two decades. Experimental work by the Wüthrich group is currently pursued primarily at The Scripps Research Institute in California (USA) and at ShanghaiTech University in China. Read more about our research.